William G. Kaelin Jr., Sir Peter J. Ratcliffe and Gregg L. Semenza were awarded the 2019 Nobel Prize in Physiology or Medicine for their discoveries of how cells sense and adapt to oxygen availability.
Kaelin and Ratcliffe’s labs focused their efforts on the transcription factor HIF (hypoxia-inducible factor). This transcription factor is critical in the cellular adaptation of to changes in oxygen availability.
When oxygen levels are elevated cells contain very little HIF. Ubiquitin is added to the HIF protein via the VHL complex and it is degraded in the proteasome. When oxygen levels are low (hypoxia) the amount of HIF increases.
In 2001 both groups published articles characterizing the interaction between VHL and HIF, and these articles were referenced by the Nobel Prize Organization in their press release about this year’s award. (1,2). Both studies demonstrated that under the normal oxygen conditions hydroxylation of proline residue P564 enabled VHL to recognize and bind to HIF.
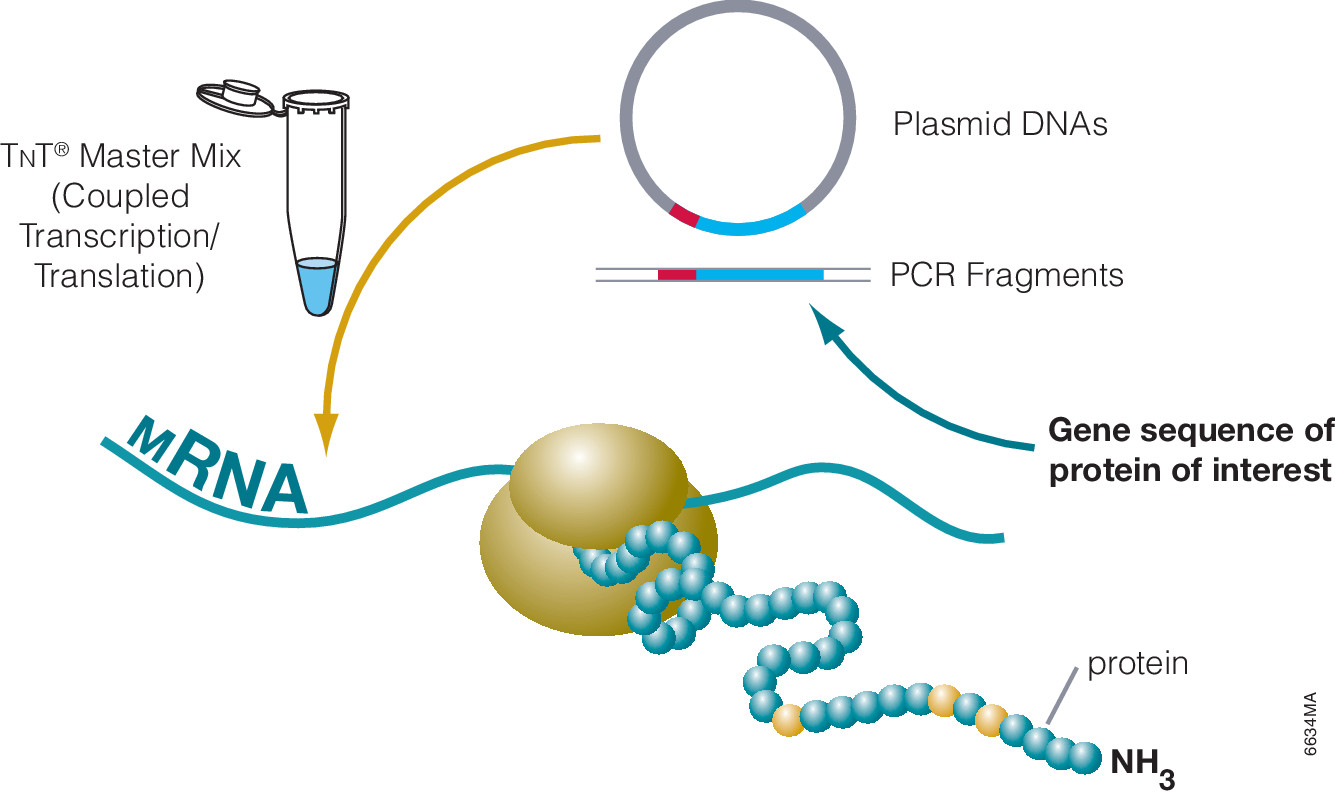
The use of cell free expression (i.e., TNT Coupled Transcription/Translation System) by both labs was key in the characterization of the VHL:HIF interaction The labs utilized HIF and VHL 35-S labeled proteins generated via the TNT system under both normal or in a hypoxic work station to:
- Determine the affect of ferrous chloride and cobaltous chloride on the interaction
- Map the specific region of HIF required for the interaction to occur (556-574)
- Determine the effect of HIF point mutations on the interaction
- Use synthetic peptides to block the interaction
- Conclude that a factor in mammalian cells was necessary for the interaction to occur.
Literature Cited
- Ivan, M et al. (2001) HIF Targeted for VHL-Mediated Destruction by Proline Hydroxylation: Implications for O2 Sensing. Science 292: 464–67.
- Jaakkola, P. et al. (2001) Targeting of HIF-α to the von Hippel-Lindau Ubiquitylation of Complex by O2– Regulated Prolyl Hydroxylation. Science 202, 468–72 .