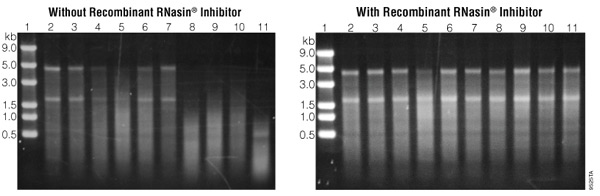
RNase, back in the early 1990s, posed a serious threat to laboratories working with RNA isolation. My graduate work involved isolating RNA from the tissues of Lyme disease-infected mice and hamsters. We struggled to DEPC-treat glass and plasticware, or autoclave anything that could be autoclaved, kept tissues cold during RNA harvest and held our breaths (truly, as aerosol could be another source of ribonuclease) until PAGE proved us successful in RNA isolation.
Ribonuclease (RNase) was omnipresent and the arch rival of our work, across several species, due to its RNA destroying abilities.
Now, a July 13, 2015 publication by researchers at the University of Wisconsin-Madison provided both a catch-up for this former lab rat on modern day research with and knowledge of RNase, as well as an exciting look at what may be a real purpose for this RNA-destroying molecule: RNase has moved to clinical trials due to the discovery of it’s cytotoxicity for cancer cells.
Raines’ group in the Department of Chemistry at UWI-Madison published in ACS Central Science their findings on the ligand that RNase 1 uses to attach to human cancer cells, in the article, “Human Cancer Antigen Globo H is a Cell-Surface Ligand for Human Ribonuclease 1”. Continue reading “A Reason for Ribonuclease: From Laboratory Nuisance to Cancer Therapeutic”