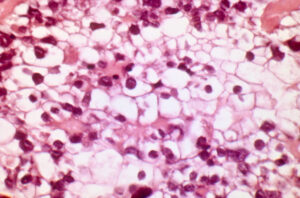
In a 2010 study of ovarian clear cell carcinoma , Jones and colleagues report a discovery of a gene that is mutated in high frequency and is involved in epigenetic regulation of gene expression.
Ovarian Clear Cell Carcinoma (OCCC) is one of the most treatment-resistant, aggressive types of ovarian cancer, and it typically progresses stepwise from endometriosis to malignancy. Mutations in the phosphotidylinositol-3-kinase gene and amplification of a region of the long arm of chromosome 20, which is also associated with breast cancer, are the two genetic aberrations most commonly associated with OCCC.
In this latest study, the authors sequenced the coding regions of eight tumor genomes. They identified four genes that were mutated in at least two of the tumors. They then sequenced these four genes in tumor and normal tissues from an additional 34 OCCC patients. Two of the four genes were disrupted in a significant percentage of the total of 42 cases examined. The PIK3CA gene was disrupted in 40% of the tumors, and the ARID1A gene was mutated in 57% of the tumors.
The PIK3CA gene is an oncogene that is involved in signal transduction, and the mutations identified in this gene were restricted to a few codons. Alterations in this gene that have been functionally characterized are activating mutations, typical for oncogenes, in which aberrant or excessive activity leads to proliferation or unrestricted growth.
The mutations in the ARID1A gene were predicted to truncate the protein either through introducing premature stop codons or creating out-of-frame insertions or deletions. Such mutations would presumably reduce or eliminate protein function, and the cancer phenotype would be associated with the loss of function of a gene that is normally a tumor suppressor.
The ARID1A gene is particularly interesting because it encodes a component of the ATP-dependent chromatin-remodeling complex SWI/SNF. This complex is involved in chromatin reorganization during gene activation and presumably regulates access to DNA by changing chromatin structure. Such function makes it part of the epigenetic regulation of the genome.
Epigenetic regulation of gene expression is heritable, and refers to changes in gene activity that do not involve alterations to the primary DNA sequence, but rather modifications to the DNA molecule itself or to the chromatin structure.
Identifying a gene involved in chromatin/nucleosome structure as a major player in cancer is important, because although epigenetic changes are prevalent in many cancer types, the relationship between the genetic changes and the epigenetic profiles of tumors is not well understood. The identification of genes directly involved in chromatin remodeling, histone modification and other epigenetic events may be key to understanding which epigenetic changes are important for tumorigeneis and, potentially, may provide an entirely new set of pathways for therapeutic targets.
Literature Reviewed
Jones, S., Wang, T., Shih, I., Mao, T., Nakayama, K., Roden, R., Glas, R., Slamon, D., Diaz, L., Vogelstein, B., Kinzler, K., Velculescu, V., and Papadopoulos, N. (2010). Frequent Mutations of Chromatin Remodeling Gene ARID1A in Ovarian Clear Cell Carcinoma Science DOI: 10.1126/science.1196333
Michele Arduengo
Latest posts by Michele Arduengo (see all)
- The Casual Catalyst: Science Conversations and Cafes - July 18, 2024
- Cancer Moonshot: Solving Tough Problems - May 28, 2024
- Automated Sampling and Detection of ToBRFV: An Emerging Tomato Virus - April 25, 2024
