Researchers having been sharing plasmids ever since there were plasmids to share. Back when I was in the lab, if you read a paper and saw an interesting construct you wished to use, you could either make it yourself or you could “clone by phone”. One of my professors was excellent at phone cloning with labs around the world and had specific strategies and tactics for getting the plasmids he wanted. Addgene makes this so much easier to share your constructs from lab to lab. Promega supports the Addgene mission statement: Accelerate research and discovery by improving access to useful research materials and information. Many of our technology platforms like HaloTag® Fusion Protein, codon-optimized Firefly luciferase genes (e.g., luc2), and NanoLuc® Luciferase are present in the repository. We encourage people to go to Addgene to get new innovative tools. Afterall, isn’t science better when we share?
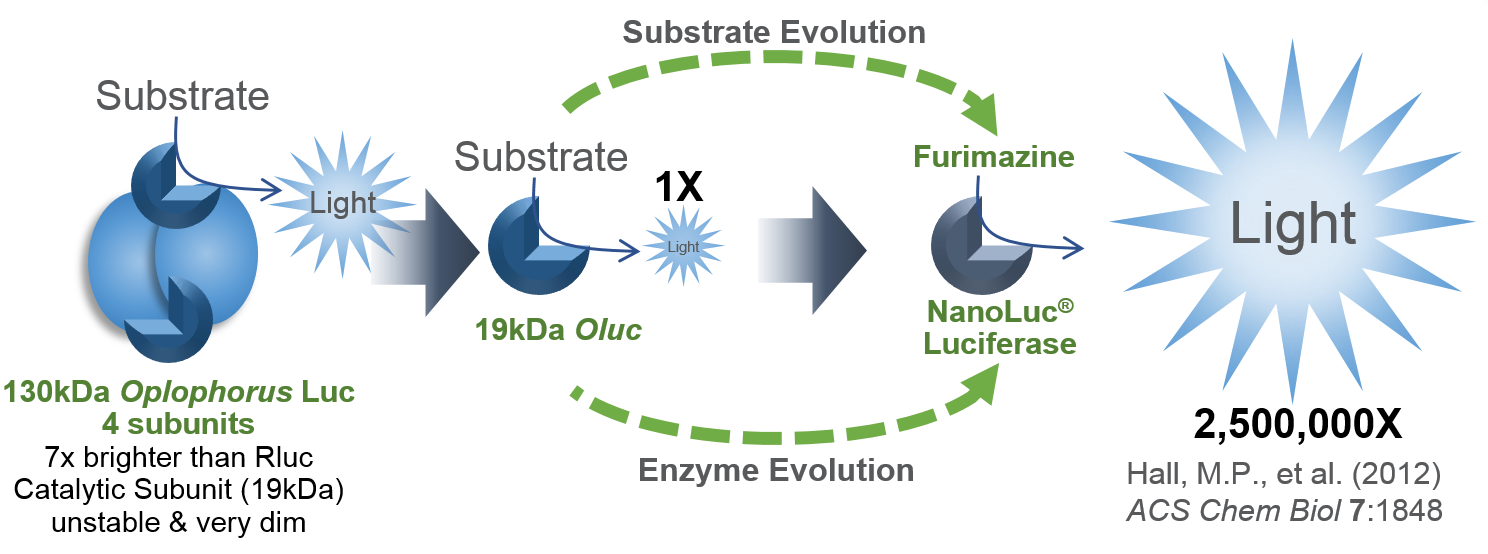
I’d like to focus on some tools in the Addgene collection based on NanoLuc® Luciferase (NLuc). The creation of NanoLuc® Luciferase and the optimal substrate furimazine is a good story (1). From a deep sea shrimp to a compact powerhouse of bioluminescence, NLuc is 100-fold brighter than our more common luciferases like firefly (FLuc) and Renilla (RLuc) luciferase. This is important not so much for how bright you can make a reaction but for how sensitive you can make a reaction. NLuc requires 100-fold less protein to produce the same amount of light from a Fluc or RLuc reaction. NLuc lets you work at physiological concentrations. NLuc is bright enough to detect endogenous tagged genes generated through the CRISPR/Cas9 knock-in. NLuc is very inviting for endogenous tagging as it is only 19kDa. An example is the CRISPaint-NLuc construct (Plasmid #67178) for use in the system outlined in Schmid-Burgk, J.L. et al (2).
Two applications of NanoLuc® Technology have caught my attention through coupling the luciferase with fluorescent proteins to make better imaging reporters and biosensors. Continue reading “Shining Stars: Cool NanoLuc® Plasmid Constructs Available Through the Addgene Repository”


